DNA collected from environmental samples reveals the diversity of extant mammals at Jasper Ridge
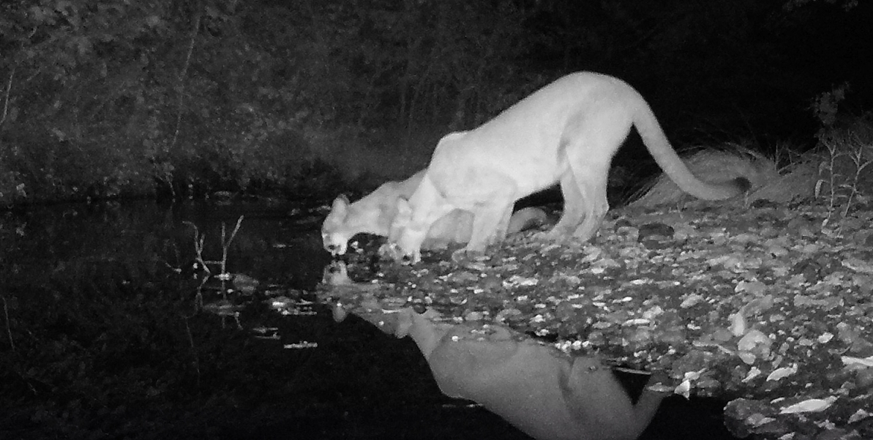

Kevin Leempoel, Trevor Hebert, Elizabeth A. Hadly
Note: Dr. Kevin Leempoel will be presenting his latest research during our Jasper Ridge Evening Lecture series on Tuesday, March 17 at 4 PM.
In conservation, faunal diversity surveys are essential to document eventual changes in diversity and abundance over time. They are, however, time-consuming, expensive, and usually detect a limited number of species. As an alternative to existing survey methods, conservation biologists are exploring Environmental DNA (eDNA), the collection of DNA from samples such as dirt or water, as one of the most promising tool for biodiversity surveys. Once DNA is extracted from the sample, it is sequenced and compared to databases in order to find out to which species these DNA fragments belong to. While this may sound rather straightforward, many unknowns and issues are preventing eDNA from going mainstream. For example, we do not know how long the DNA stays detectable in the environment or how frequent a species must be to be detected. Most importantly, comparison with existing approaches are necessary, yet is lacking for terrestrial mammals.
In our study "A comparison of eDNA to camera trapping for assessment of terrestrial mammal diversity", we compared the detection of terrestrial mammals using eDNA analysis of soil samples against confirmed species observations from camera-traps. We took advantage of the ongoing long-term (~9-yr) camera-trapping study conducted here at Jasper Ridge Biological Preserve that has been showcased previously here at Stanford News and Stanford Magazine. In this study, we found that all mammals regularly recorded with cameras were detected in eDNA (Fig. 1).
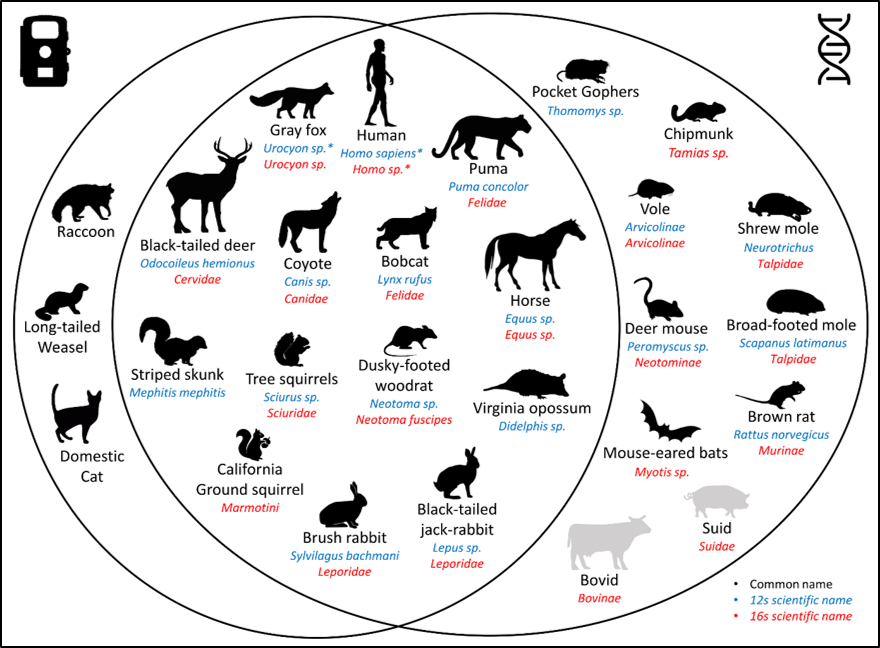
Figure 1. Venn Diagram showing species recorded by camera traps and species detected with eDNA. Mammal DNA was sequenced for two fragments (called metabarcodes) of different sizes. The fragment in the mitochondrial gene 16s is 70 base pairs long and the fragment in the gene 12s is 200 bp. Species detected with the M16s are marked in red and those detected with the M12s are marked in blue. Scientific names are given at the maximum rank reached with eDNA. Species known to be present in the study area are in black. Species absent from the study area but detected with eDNA are in grey. Species considered as contaminant are indicated with an asterisk (*).
In addition, eDNA reported many unrecorded small mammals whose presence in the study area is otherwise documented (Fig. 2). We also found that a longer DNA fragment achieved a similar efficiency as a shorter one, enabling a higher taxonomic resolution, which means we were able to identify species from these DNA fragments, as opposed to genus or family only (see examples of the wildlife captured . These encouraging results support that eDNA-based monitoring should become a valuable part of ecosystem surveys. Unfortunately, we also report that most mammals are missing from reference databases, therefore hindering the applicability of this novel technology worldwide.
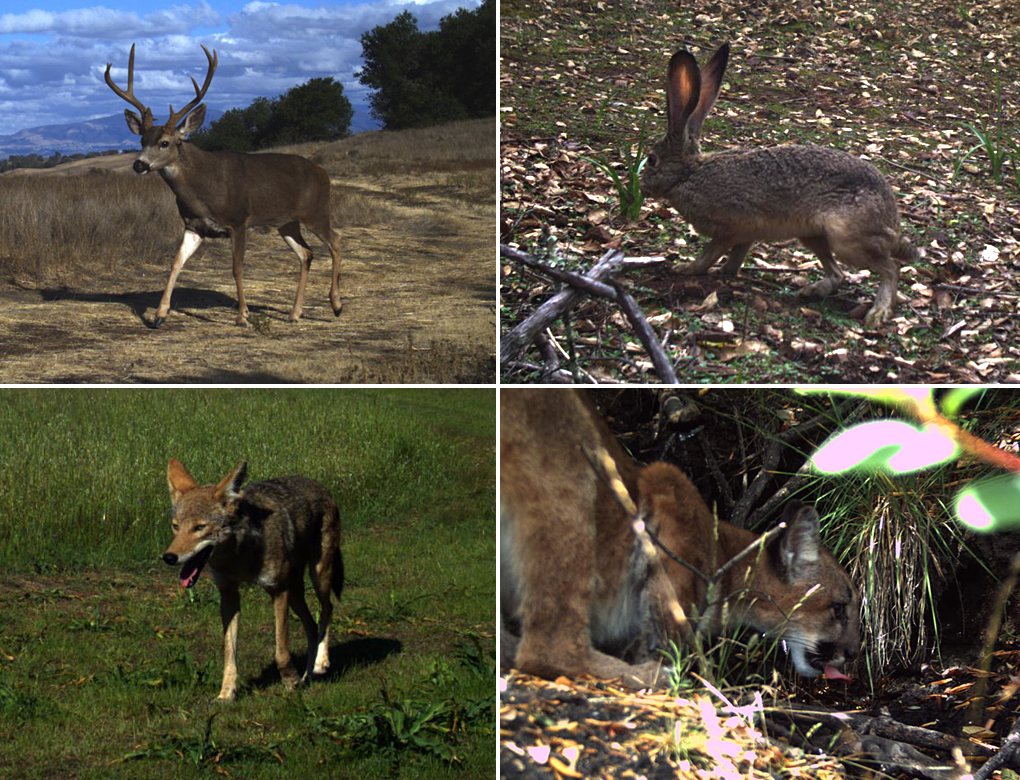 Figure 2. Images of Jasper Ridge wildlife from camera traps used in this study (clockwise from top left): Black-tailed deer (Odocoileus hemionus), black-tailed jackrabbit (Lepus californicus), mountain lion (Puma concolor) and coyote (Canis latrans).
Figure 2. Images of Jasper Ridge wildlife from camera traps used in this study (clockwise from top left): Black-tailed deer (Odocoileus hemionus), black-tailed jackrabbit (Lepus californicus), mountain lion (Puma concolor) and coyote (Canis latrans).
Link to article: A comparison of eDNA to camera trapping for assessment of terrestrial mammal diversity
 Dr. Kevin Leempoel, Postdoctoral researcher
Dr. Kevin Leempoel, Postdoctoral researcher
Jasper Ridge Biological Preserve, Hadly Lab
Stanford University